The Nishiguchi Laboratory
Philosophy of Research Program
Vibrio light organ association provides an experimentally tractable model system for the study of the mechanisms underlying bacterial colonization of animal tissues, and exploring the population dynamics, ecology and community dynamics between squid hosts and their bacterial symbionts. In this animal-bacterial system, there is no contact between host and symbiont throughout embryogenesis; the symbiosis is established within hours (environmentally) after the juvenile squids are hatched into the surrounding environment. During this infection period, specific strains of Vibrio from the water column are able to recognize and colonize the sterile light organs of the newly hatched juveniles. Bacteria that are capable of persisting in the developing light organ after this infection period represent a successful association by one specific subset (or subspecies) of Vibrio strains. This specificity is maintained throughout the squid’s life history; however, other symbiotic strains of Vibrio isolated from different host squid’s are able to cross-infect juveniles of closely related species. Competition experiments have proven that there is a hierarchy among these Vibrio strains that is congruent to the phylogenetic relationships of the host squids, and that some level of molecular recognition or specificity is involved in the establishment of each sepiolid species partnership. This provides the basis for further investigation of the molecular differences between closely related strains and species of competent vibrios and whether these differences are the result of the complex interactions (both biotic and abiotic) that have led to the evolution and radiation of several species of Vibrio bacteria.
The following paragraphs are summaries of some of the ongoing research projects in my laboratory.
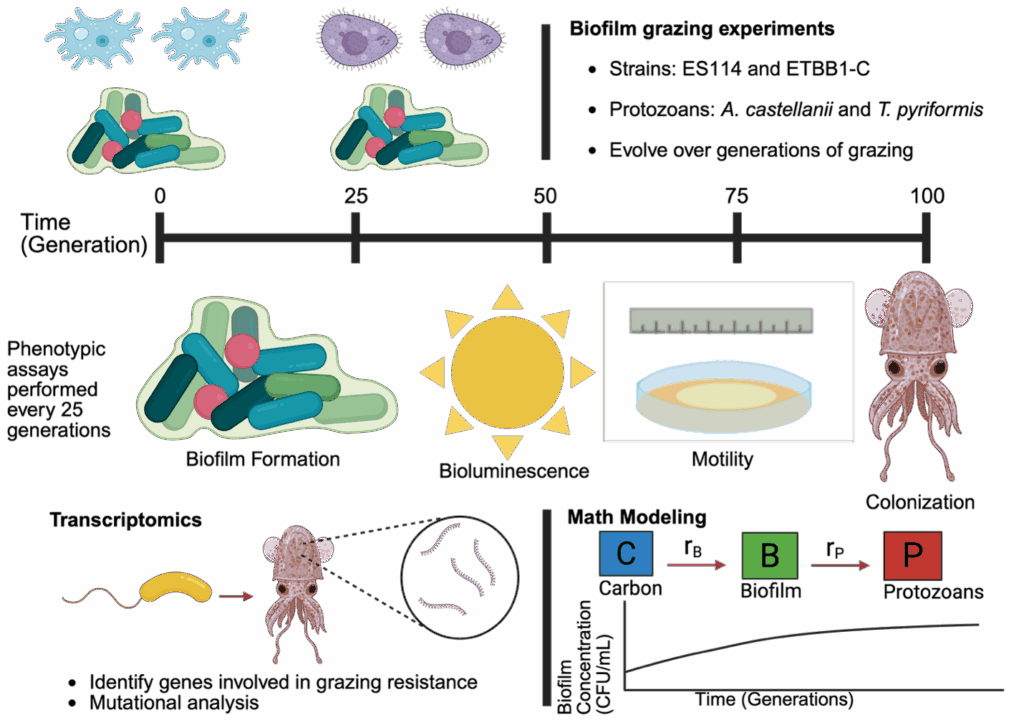
Biofilm Grazing: Predation Impacts on Host-Microbial Symbiosis
Biofilms are a community of cells adhered to a surface. Biofilm-like aggregates play an important role in colonization their squid host. When outside of the host, they are susceptible to ecological stressors such as protozoan predation. This project focuses on V. fischeri biofilms that are evolved under protozoan predation. It is hypothesized that predation will lead to changes in biofilm formation, which in turn will impact their colonization fitness in the squid host. These changes are determined through various fitness characteristics such as biofilm, bioluminescence, and motility. Ancestral and evolved strains can be compared through colonization assays in the host squid to determine if there are trade-offs between factors influencing symbionts outside the squid compared to host selection inside the squid. Differences resulting from long term predation will be ascertained through dual RNA-seq analysis of both squid and symbiont transcriptomes. Lastly, math models are being developed to better understand the factors involved in biofilm response to protozoan predation and their downstream effects on symbiosis. Results from this work will help inform the importance of predation as an ecological stressor on host-microbial symbiosis.
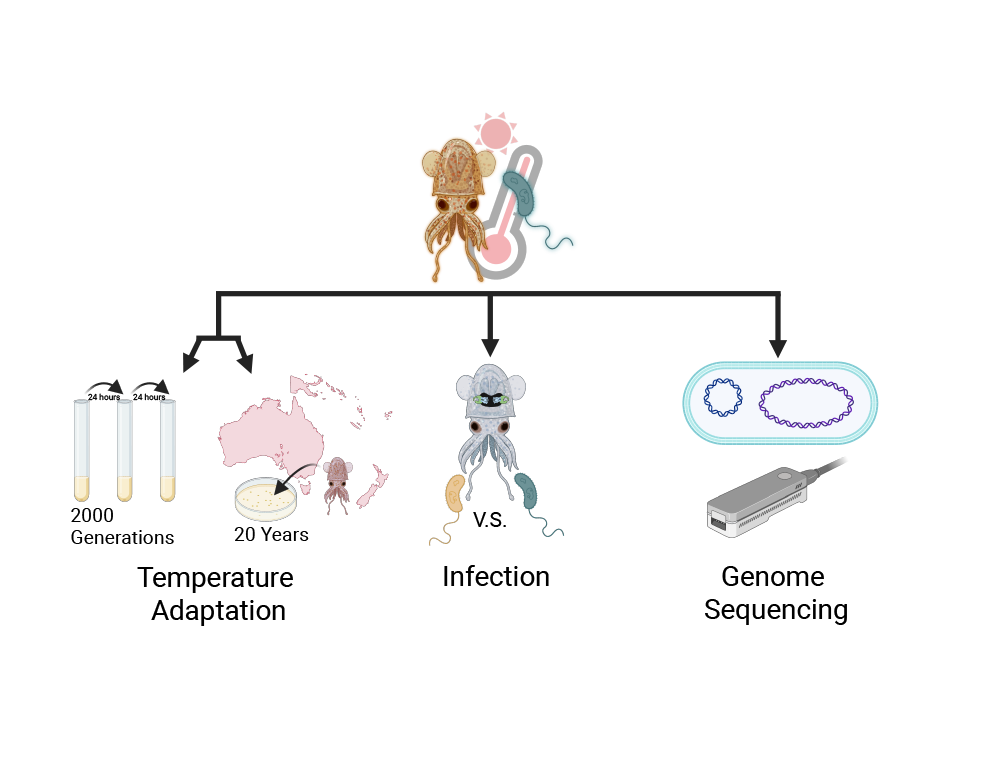
Long term experimental evolution of symbiotic V. fischeri under temperature duress
Stable microbial associations contribute to host fitness, but recent anthropogenic warming is inducing stress on individuals and their host communities. Examining how symbiotic bacteria respond to such changes can provide insight into how their hosts accommodate climate stress. The beneficial association between bobtail squids (Cephalopoda: Sepiolidae) and their Vibrio symbionts is an ideal model to understand how microbial associations can be used as a yardstick to measure adaptation to climate change. To study the impact of thermal stress over time we have taken multiple isolates of V. fischeri and adapted them to experimental temperatures by culturing them constantly for 2,000 bacterial generations. Additionally, our lab is revisiting field locations which were sampled twenty years previously and isolating symbiotic Vibrio from squids. The field-isolated and laboratory-grown samples will be studied for alterations in critical symbiosis- relevant phenotypes, mutations in their genome, and changes in host infection proficiency. The combination of field sampling and laboratory techniques will provide multiple perspectives on the impact of elevated temperatures on symbiont function. Together, this will elucidated behaviors of a symbiotic relationship under increasing temperature stress.
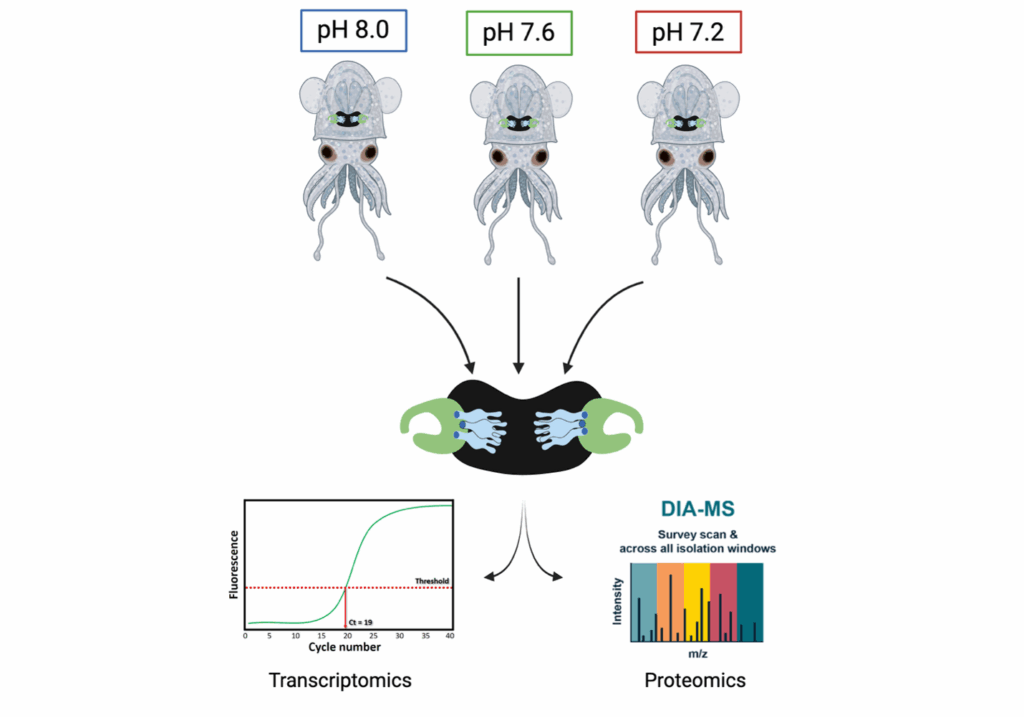
Long term experimental evolution to assess environmental changes in ocean acidification (pH).
Due to the broad temperature gradient over a wide geographic range, E. tasmanica is believed to be more robust and is therefore an excellent model for assessing how environmental stressors will affect the symbiotic interactions within the light organ. Given the abiotic changes that occur with environmental variation (pH, temperature) our lab is determining whether lower pH (higher acidity) affects the symbiosis between E. tasmanica and their bioluminescent symbiont V. fischeri by analyzing gene and protein expression from the light organs of juvenile squids raised under various pH scenarios. We am characterizing which genes are positively expressed and by doing so, determining pathways involved in regulating changes while identifying transcription factors that regulate symbiosis in response to lower pH. We plan to identify proteins and protein families that have low pH stability to understand how this symbiosis changes at the molecular scale in response to higher acidity. This work will help us better understand the immediate impact of low pH due to the changing pressures that occur with climate crises.
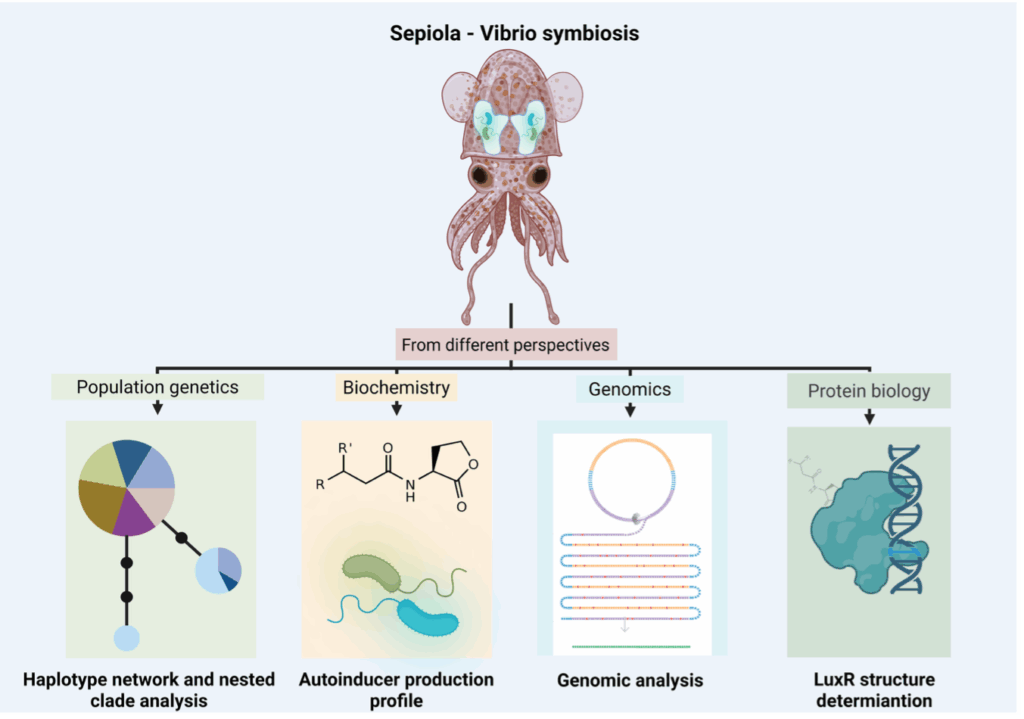
Variation in Vibrio symbionts between Mediterranean and Atlantic sepiolid squids
This project focuses on the genus Sepiola, native to the Mediterranean Sea and the Atlantic Ocean, and its bacterial symbionts: Vibrio fischeri, V. logei, and V. wodanis. Unlike the widely studied Euprymna scolopes–Vibrio fischeri model from the Indo-Pacific, Sepiola species host multiple Vibrio species simultaneously within their light organ, coexisting in both time and space. This difference provides a unique window to explore the Sepiola–Vibrio association and determine whether these bacteria behave differently when sharing the same host. We are assessing whethe these Vibrio strains cooperate to colonize and thrive in the squid’s light organ, or whether they are competing for space and resources. We have examined the geographical barriers such as ocean currents and how they influence the genetic structure of both the host (Sepiola spp.) and its bacterial symbionts. Subsequently, we are combining a comparative genomics approach to biochemical and structural analyses, to explore the role of the LuxR protein to determining whether bacterial communication among the symbiont community works cooperatively. This includes studying the interaction between LuxR and acyl-homoserine lactones (AHLs), the quorum-sensing molecules bacteria use to coordinate behaviors such as bioluminescence, which is essential for the symbiosis. We hope to clarify whether coexisting Vibrio species cooperate or compete and offer insight into how bacterial communication influences beneficial associations.
Vibrio fischeri metabolomics during symbiosis with sepiolid squids
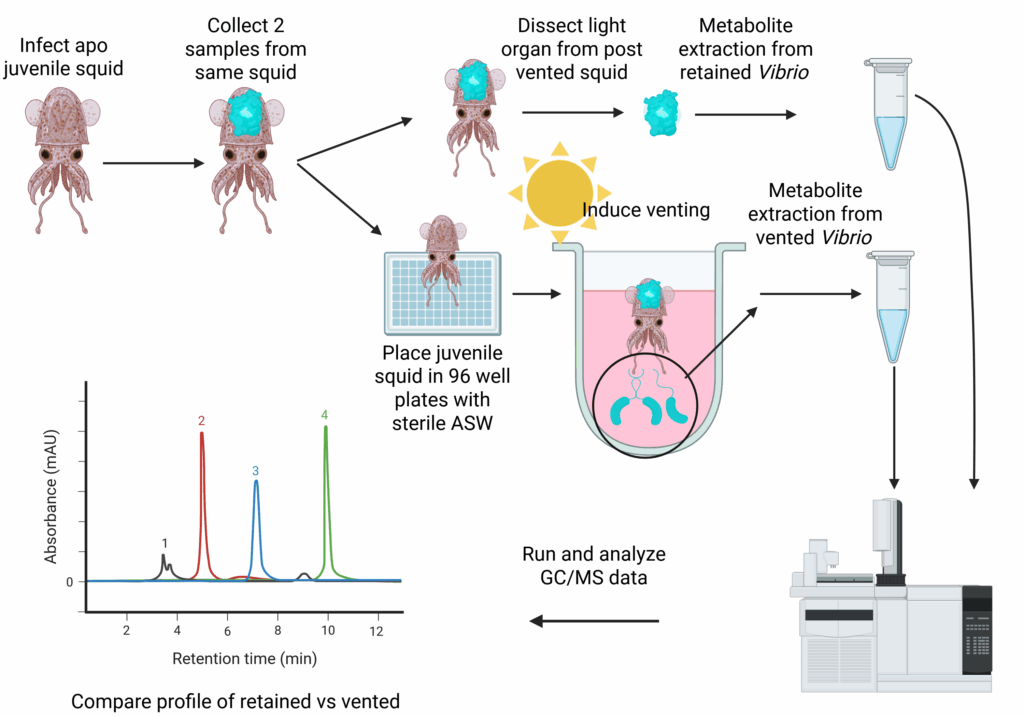
This project focuses on how Vibrio fischeri metabolism is shaped by environmental and host- associated states. On the left V. fischeri exist in the free-living state in seawater. It then encounters its host (Euprymna scolopes) and undergoes metabolic shifts as is colonizes the light organ crypts and adapts to host-derived nutrients and signals. Within the light organ crypts, V. fischeri exhibits specialized metabolic profiles optimized for symbiosis. Each morning most of the bacteria in the light organ are vented back into the environment allowing the bacteria that are retained to repopulate the light organ. This venting behavior provides an opportunity to compare host-retained and released populations. Using untargeted metabolomics and RNA-Seq, we plan to investigate how these transitions influence bacterial metabolism, gene expression and adaptation.
 Michele K. Nishiguchi, PhD
Michele K. Nishiguchi, PhD

